In classrooms, breeding barns and research benches alike, knowing how different crosses work helps you predict trait patterns and design experiments. Whether you’re reviewing Mendel’s classic crosses or planning a selection scheme, a compact reference keeps the comparisons clear.
There are 18 Types of Genetic Crosses, ranging from Advanced Intercross Line (AIL) to Trihybrid cross; for each type you’ll find below Parental genotypes,Expected progeny ratio,Common use (max 15 words) to make comparisons fast and practical — you’ll find below the organized list and details.
How do I pick the best cross for my experiment?
Choose based on your goal: map genes (use intercrosses or AIL for recombination), test dominance (use monohybrid/dihybrid), or combine traits (use trihybrid or backcross). Match your population size and expected ratios to the statistical power you need.
What progeny ratios should I expect from common crosses?
Classic crosses give predictable Mendelian ratios (e.g., 3:1 for monohybrid, 9:3:3:1 for dihybrid), but linkage, epistasis or incomplete dominance change those outcomes — always check parental genotypes and assumptions before interpreting results.
Types of Genetic Crosses
| Name | Parental genotypes | Expected progeny ratio | Common use (max 15 words) |
|---|---|---|---|
| Monohybrid cross | AA x aa or Aa x Aa | Phenotypic 3:1; genotypic 1:2:1 | Test single-gene dominance |
| Testcross | Aa x aa | Phenotypic 1:1 if heterozygote tested | Determine dominance/heterozygosity |
| Dihybrid cross | AaBb x AaBb | Phenotypic 9:3:3:1 | Study independent assortment |
| Trihybrid cross | AaBbCc x AaBbCc | Phenotypic 27:9:9:9:3:3:3:1 | Complex multi-gene inheritance |
| Backcross | F1 x parental (e.g., Aa x AA or Aa x aa) | 1:1 when backcrossed to recessive parent | Introgression; confirm genotypes |
| Reciprocal cross | AA x aa and aa x AA | Same ratios; detects maternal effects | Test maternal/cytoplasmic effects |
| Selfing | Aa (self) or plant self-fertilization | Genotypic 1:2:1; phenotypic 3:1 | Fix traits; produce inbred lines |
| Outcross | Two unrelated individuals (Aa x Bb typical) | Varies; increases heterozygosity | Introduce variation; hybrid vigor |
| F1 cross | F1 x F1 (Aa x Aa) | Genotypic 1:2:1; phenotypic 3:1 in F2 | Produce F2 segregation; study dominance |
| Complementation cross | mutA x mutB (two mutants) | F1 wild-type or mutant (qualitative) | Test if mutations allelic |
| Diallel cross | All parents crossed in all combinations | Varies; many combinations | Estimate combining ability in breeding |
| Factorial cross | Set A parents x Set B parents (e.g., A1xB1) | Varies by combination | Test interactions; breeding combinations |
| Three-way cross | (A x B) x C | Varies; hybrid combining multiple parents | Create hybrids combining three parents |
| Double cross | (A x B) and (C x D) then (AB) x (CD) | Varies; broad hybrid vigor | Develop broad-based hybrids |
| Introgression (recurrent backcrossing) | Donor allele Aa x recurrent AA, repeated BCs | Donor allele fraction reduces each backcross | Transfer single trait into elite background |
| Recombinant Inbred Line (RIL) cross | P1 x P2 then many selfing generations | Lines fixed; parental allele ~50% across lines | Create homozygous mapping populations |
| Advanced Intercross Line (AIL) | P1 x P2 then many random intercross generations | More recombination than F2; varied genotypes | Increase mapping resolution via recombination |
| North Carolina designs (I/II/III) | Structured crosses among multiple sires/dams | Varies; used for variance component estimation | Partition genetic variance components |
Images and Descriptions

Monohybrid cross
A single-gene cross, typically Aa x Aa or AA x aa, used to reveal dominance relationships. Predictable phenotypic 3:1 in F2 and genotypic 1:2:1. Use for simple Mendelian trait teaching and basic inheritance experiments.

Testcross
Cross between an individual of unknown genotype and a homozygous recessive tester. A 1:1 phenotypic split shows the tested parent was heterozygous; all uniform offspring indicate homozygosity. Widely used to reveal hidden alleles.
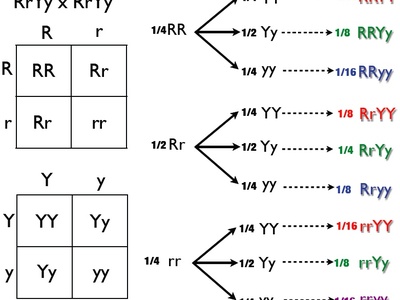
Dihybrid cross
Cross tracking two independent genes, e.g., AaBb x AaBb. Classic F2 phenotypic ratio 9:3:3:1 if genes assort independently and show dominance. Useful to test Mendel’s law of independent assortment and epistasis deviations.

Trihybrid cross
Cross involving three independently assorting genes; standard F2 gives a 27:9:9:9:3:3:3:1 phenotypic distribution assuming simple dominance. Use to illustrate multi-locus inheritance, linkage effects, and interactions among three traits. Often used in teaching and problem sets.

Backcross
Cross F1 offspring back to one parent (recurrent parent) to test allele dominance or transfer traits. Backcross to a homozygous recessive gives a 1:1 segregation. Widely used in breeding and mapping of traits.

Reciprocal cross
Performing a cross in both directions (male A x female B and vice versa) to reveal maternal inheritance, sex-linkage, or cytoplasmic effects. If F1 differs between directions, non-Mendelian maternal or sex-linked factors may be involved.

Selfing
Self-fertilization (selfing) of heterozygotes or hermaphrodites produces predictable segregation: genotypic 1:2:1 and phenotypic 3:1 for a single locus. Used to create inbred lines and reveal recessive alleles over generations. Common in plant genetics and Mendelian experiments.

Outcross
Cross between unrelated individuals or lines to combine traits and increase heterozygosity. Expected outcomes depend on parental genotypes; often used to produce hybrids with vigor, break inbreeding, or introduce new alleles into a population.

F1 cross
Cross F1 siblings or selfed F1s to generate F2 progeny. The F2 shows segregation (genotypic 1:2:1 and phenotypic 3:1 for single genes), revealing hidden variation and allowing mapping of traits segregating in offspring.

Complementation cross
Cross two recessive mutants to test whether mutations affect the same gene (allelic) or different genes (complementation). A wild-type F1 indicates complementation (different genes); a mutant F1 suggests allelism. Widely used in genetic screens.

Diallel cross
Mating every parent with every other parent (including selfs or excluding selfs) to evaluate general and specific combining abilities. Helps breeders identify parents that produce superior hybrids and partition additive versus non-additive effects.

Factorial cross
Cross two sets of parents in all possible pairings (A x B) to study genetic interactions and combine traits systematically. Useful for evaluating genotype-by-environment or combining specific parental lines in breeding programs.

Three-way cross
First cross two lines (A x B) and then cross their hybrid to a third parent C. Used to combine desirable traits from three sources and produce commercial hybrids with balanced performance and heterosis.

Double cross
Make two initial crosses (A x B and C x D), then cross their F1 hybrids to each other. Produces genetically diverse hybrids that can combine favorable alleles from four parents and often show strong heterosis across traits.

Introgression (recurrent backcrossing)
Repeatedly backcross a donor carrying a trait into an elite recurrent parent to move a specific allele while restoring the recurrent genome. Donor genome fraction halves each generation, enabling precise trait transfer with minimal unwanted background.

Recombinant Inbred Line (RIL) cross
Cross two inbred parents and repeatedly self progeny (often >6 generations) to produce RILs. Lines become homozygous and capture recombination breakpoints, making them ideal for high-resolution mapping and replicable genetic studies. Widely used in plant and animal genetics.

Advanced Intercross Line (AIL)
Initiate from two parents and perform several generations of random intercrossing before inbreeding. Additional meioses increase recombination breakpoints, improving mapping resolution for QTLs compared with standard F2 or early generation RILs. Used in advanced genetic mapping studies.

North Carolina designs (I/II/III)
Sets of structured mating designs (North Carolina I, II, III) that cross groups of males and females in specific patterns. Used to estimate additive and dominance variance components and inform breeding value and inheritance structure.